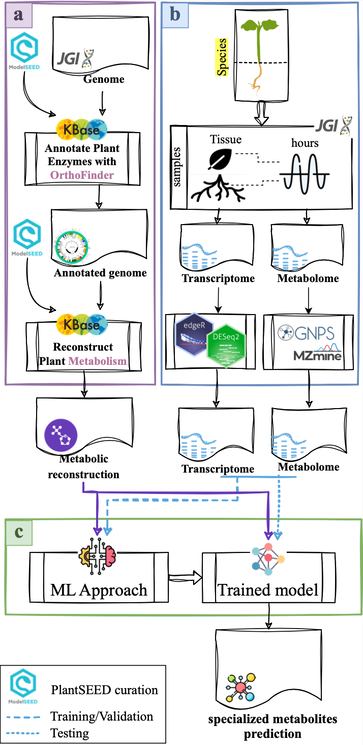
Resolving the spatial dynamics of single cell circadian regulatory networks
Internal circadian oscillators are common to all organisms from cyanobacteria to humans. This endogenous ~24h timekeeper synchronizes metabolic and physiological processes with the environment to maintain fitness. The circadian oscillator consists of a series of transcriptional feedback loops that provide diel regulation of gene expression. In plants, we have a good understanding of the molecular players comprising the circadian oscillator; however, this knowledge is based on whole tissue or mixed tissue samples. We now know that circadian regulation of transcription is highly dependent on cell type. How does clock regulation at the cell level lead to coordinated tissue responses and what alterations to the gene regulatory network (GRN) facilitate these signals?

NIGMS 1R35GM146837-01
PI: Kathleen Greenham
|
Enhancing stress tolerance using a synthetic phase-dependent stress response.
Circadian biologists have associated large portions of the transcriptome with temporal regulation by the circadian clock. One consequence of this internal timekeeping in plants is time of day dependent responses to abiotic and biotic stress. Given the energy costs associated with mounting a defense response it is not surprising that plants restrict the response to the time when its effectiveness is maximal. Rather than thinking of stress response as simply on or off, we focus on fine-tuning the timing of the response in a more 'turn up or down' approach. Adjusting the amplitude of the response within the normal phase domain and maintaining the timing of essential growth processes has the potential to enhance stress tolerance without a reduction in fitness. |